BY RANDY MCINTOSH
Data I know thee,
I know thy face,
I have seen the fear in thine eyes,
I shall meet thee on the field of battle,
And I shall conquer thee...
As part of our blog series, we will occasionally post summaries of our lab meetings. This blog will be the first of those, where I presented some recent observations on quality control of data pipelines for neuroimaging data.
We are in the midst a large project using TheVirtualBrain (TVB) to model dementia, with the hope of characterizing subtypes early in the disease progression based on the biophysical parameters we get from TVB. We’ve had success in doing this in stroke recovery and our colleagues in Marseille are finding great traction in epilepsy for pre-surgical planning.
Since I have a lot of free time now, I elected to help out with the data processing, which relies on publicly available dataset in ADNI and PPMI. Given the extensive atrophy that we’ve seen in aging and dementia, we opted to put in several quality control steps to make sure that any artefacts were minimal. In TVB, we use functional connectivity (correlation of time series of brain data) as the target to optimize the model, so if the data have artefact, that can affect the correlations, which of course can render the model invalid.
We noted, to no surprise, that data from patients are messy. The varying levels of atrophy, pervasive head-motion artefact, and misalignment of structural and functional data can interact in weird and wonderful ways that wreck havoc on data quality, as I show in the example below.
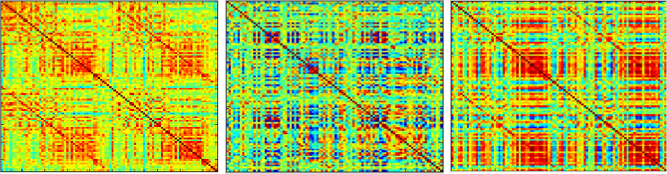
These are three correlation matrices (aka functional connectivity matrices) from these subjects with Alzheimer’s disease from the ADNI database. The correlations, scaled from -1 (blue) to 1 (red), are from resting-state fMRI data expressed as time-series from a region of interest template. What is clear are huge differences across subjects. The leftmost matrix shows the “typical” features in a functional connectivity matrix with a moderate range of values, mostly positive, with a noticeable off-diagonal element for cross-hemispheric correlations of homologous regions (e.g., left and right primary motor cortex). The middle matrix is from a subject with severe atrophy and excess head motion, and shows quite a different pattern of correlations. The extreme range and banding pattern is usually indicative of excess artefact. The rightmost matrix is from a subject who has comparable data features as the person on the far left in terms of atrophy and head motion, but several of the regions of interest were misaligned, leading to “partial volume” artefact, where the average of a region contains values from more than just gray matter (usually CSF). The problem is that the banding pattern in the matrix occurs for some regions where the template match was fine, which suggests a either a different source of artefact, or that this is indeed that person’s functional connectivity pattern.
None of this is new in neuroimaging. The challenge is finding a way to balance rigorous checks for data quality with pushing a lot of data through. In the Halcion days of neuroimaging, with positron emission tomography and small sample sizes, you could track the pipeline step by step for any errors. This didn’t eliminate artefact, as there were examples of missed artefacts getting passed off as reliable results (you know who you are). With the era of Big Data firmly upon us, the challenges for ensuring data quality are even greater. Having a ton of data doesn’t absolve you of checking your data processing stream. In fact, its probably even more critical now as with Big Data comes a serious potential for Big Errors (see here for a nice discussion of this). My colleague, Stephen Strother, and his group have advocated an optimization approach for pipelines, which modifies the parameters of a processing pipeline in favour of reproducibility within subject. It takes a bit longer to run, but given the amount of effort spent collecting data, and then analyzing, the few extra hours spent on ensuring high data quality is easy to justify.
So, yeah, with the processing power and cool tools we have available to us, we are well-prepared to do battle with our data, with ground breaking papers as evidence of our conquest.
To finish off the opening stanza:
And I shall conquer thee…
But first let me do a quick QC check :)